Molecular de-extinction could offer avenues for drug discovery by reintroducing bioactive molecules that are no longer encoded by living organisms.
Human genomes and the genomes of our ancient ancestors express proteins with natural antimicrobial properties.
Molecular de-extinction hypothesizes that these molecules could be prime candidates for safe new drugs.
Naturally produced and selected through evolution, these molecules offer promising advantages over molecular discovery using AI alone.
“We need to think big in antibiotics research,” said Dr. Cesar de la Fuente, a researcher at the University of Pennsylvania.
“Over one million people die every year from drug-resistant infections, and this is predicted to reach 10 million by 2050.”
“There hasn’t been a truly new class of antibiotics in decades, and there are so few of us tackling this issue that we need to be thinking about more than just new drugs. We need new frameworks.”
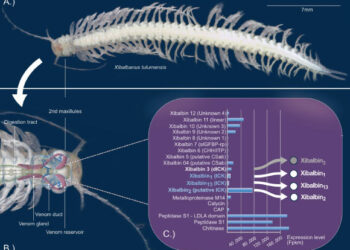
In the study, the authors explored the proteomic expressions of two extinct hominins — Neanderthals and Denisovans — and found dozens of small protein sequences with antibiotic qualities.
Their lab then worked to synthesize these molecules, bringing these long-since-vanished chemistries back to life.
“The computer gives us a sequence of amino acids,” Dr. de la Fuente said.
“These are the building blocks of a peptide, a small protein. Then we can make these molecules using a method called ‘solid-phase chemical synthesis’.”
“We translate the recipe of amino acids into an actual molecule and then build it.”
The researchers next applied these molecules to pathogens in a dish and in mice to test the veracity and efficacy of their computational predictions.
“The ones that worked, worked quite well,” Dr. de la Fuente said.
“In two cases, the peptides were comparable — if not better — than the standard of care.”
“The ones that didn’t work helped us learn what needed to be improved in our AI…
Read the full article here