Researchers at the University of Connecticut have published the first high-quality reference genome for the butternut (Juglans cinerea), a member of the walnut family Juglandaceae.
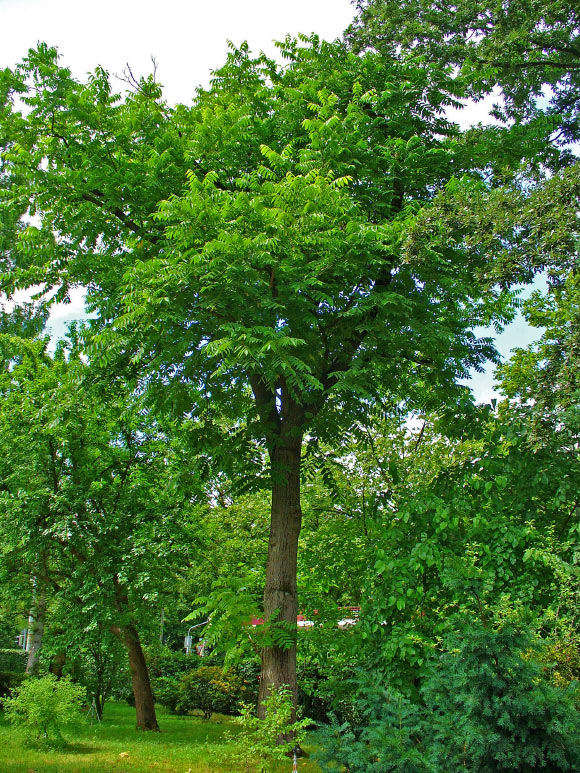
Butternut, also known as white walnut, is a deciduous, outcrossing, and wind-pollinated canopy tree native to eastern North America.
Once valued for its wood and edible nuts, today, it is currently listed as Endangered on the IUCN Red List due to decline from butternut canker, a disease caused by the fungus Ophiognomonia clavigignenti-juglandacearum.
Discovered in the 1960s and now found throughout butternut’s range, the disease had already killed more than 80% of butternut trees in numerous states by the 1990s and is threatening the species with extinction.
“We’re interested in knowing how much of the butternut population is already hybridized with the Japanese walnut (Juglans ailantifolia), and what is contributing to the genetic resistance, to the fungal infection,” said Dr. Jill Wegrzyn, a computational biologist in the Department of Ecology and Evolutionary Biology and the Biodiversity and Conservation Genomics Center at the University of Connecticut.
To address the conservation genetics gap, University of Connecticut undergraduate and graduate students worked with practitioners to generate the first reference genome for Juglans cinerea.
The chromosome-scale reference was annotated with existing transcriptomic resources and examined in an evolutionary context to recently sequenced members of the Juglandaceae family from around the globe.
“The construction of the genome was connected to a year-long training program initiated with high molecular weight extraction of the target tree,” the researchers said.
“Students worked closely with graduate students and faculty members to understand the challenges associated with genome assembly and annotation.”
“In doing so, the genome benefitted from robust methods testing,” they added.
“Finally, the program…
Read the full article here