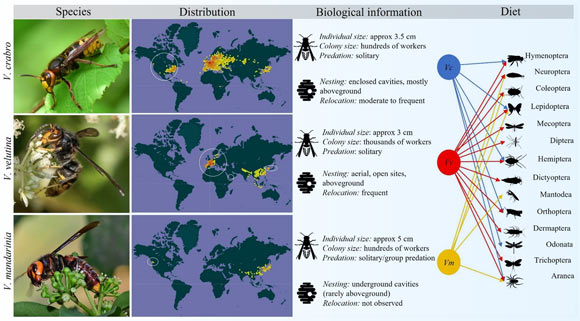
Hornets are the largest of the social wasps, and are important regulators of insect populations in their native ranges. These insects are also very successful as invasive species, with often devastating economic, ecological and societal effects. Understanding why they are such successful invaders is critical to managing future introductions and minimizing impact on native biodiversity. Critical to the management toolkit is a comprehensive genomic resource for these insects. In new research, scientists sequenced the genomes of two hornet species: the European hornet (Vespa crabro) and the Asian hornet (Vespa velutina). They also compared these genomes with those of other social Hymenoptera, including the northern giant hornet (Vespa mandarinia). The three hornet genomes show evidence of selection pressure on genes associated with reproduction, which might facilitate the transition into invasive ranges.
There are around 1,200 species of social wasps, including relatively well-known Vespidae yellowjackets and hornets, and lesser-known species of Stenogastrinae and Polistinae.
They display a huge variety of ecological and life-history characteristics; e.g. the colony size varies enormously, from species with less than 10 individuals in the society (e.g. as found in the Stenogastrine hover wasps) to those with tens of thousands of workers (e.g. Vespula species).
Some species have many reproductive queens (e.g. epiponine wasps, like Metapolybia) whilst others are monogynous (e.g. Vespa crabro); reproductive hierarchies can be regulated by conventions such as age or size, aggression or pheromones.
The first aculeate wasp genome was published in 2015 (Polistes canadensis), closely followed by the European paper wasp (Polistes dominula).
Currently there are genome sequences published for seven polistine wasp species and nine vespine wasps.
However, evolutionary analyses of these vespine genomes are lacking, and particularly so for the hornets, the Vespa genus.
“To…
Read the full article here